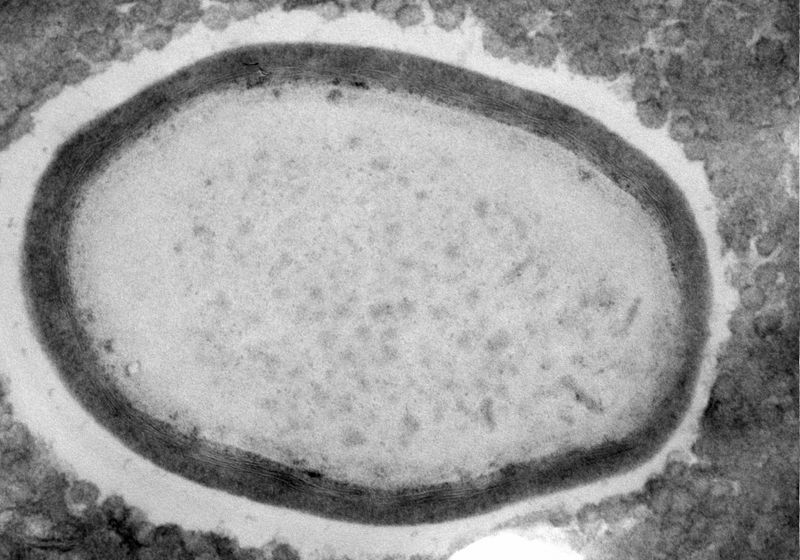
Most viruses are small and carry minimalist genomes. Even one of the largest small viruses, Vaccinia, measures merely one-fiftieth the size of a pollen grain and contains only 270 genes. Giant viruses flout these rules. With sizes that rival small bacteria and genomes that contain thousands of genes, their complexity emulates that of cellular life.
How these viruses came to be so large has been the subject of much debate. Now, scientists are finally poised to unravel the mystery of their evolutionary origins, thanks to a suite of CRISPR/Cas9-based tools described in a Nature Communications paper from January.
See “Monir Moniruzzaman Studies the Secrets of Giant Viruses”
“It was by chance that we encountered the first giant virus,” says Chantal Abergel, a virologist at Aix-Marseille University in France. “It was Mimivirus, and it was actually mistaken for a bacterium.” In the 20 years since that discovery, virologists have prioritized exploring the diversity of giant viruses. Now that they’ve found a fair few, the focus has shifted towards studying their evolution in more detail with molecular biology techniques.
Evolutionary biologists have grappled over two possible origins of giant viruses. One possibility is that they were once cellular organisms that shrunk physically and genetically over time. But most virologists now suspect giant viruses grew out of much smaller ones—though the evidence supporting either hypothesis is scant.
To begin addressing this origin question, Abergel decided to examine how the essential genes in the Pandoravirus genome are distributed. In cellular organisms, essential genes are scattered throughout the genome—so if giant viruses are essentially reduced cells, one would expect a similar pattern. Alternatively, if the genes are clumped, that could indicate the viruses’ large genomes started out in a more compact form.
One way to locate a virus’s essential genes is to knock out genes one at a time to find the ones that are needed for virus production. But to do that with a giant virus, Abergel needed a gene-editing system that worked in members of the group. With the help of Hugo Bisio, a postdoctoral researcher in Abergel’s lab, and colleagues at Aix–Marseille University, Abergel used a CRISPR/Cas9-based gene-editing system to modify the genome of the amoeba Acanthamoeba castellanii and the giant virus Pandoravirus neocaledonia, which infects it.
See “New Giant Viruses Break Records”
The CRISPR/Cas9 system was designed to delete specific genes and consists of two guide RNAs and a Cas9 scission enzyme. Similar to other CRISPR/Cas9 systems, each guide RNA contains 17 to 20 bases designed to bind to one specific location on the genome of the giant virus or the amoeba, allowing the Cas9 scission enzyme to cut the genome at that site.
See “Ten Years of CRISPR”
The amoeba A. castellanii contains 25 copies of each chromosome, making it difficult to design an efficient CRISPR/Cas9 system that could delete each gene copy. To overcome this issue, the researchers modified their CRISPR/Cas9 system to generate a chain reaction. Each time DNA was cut to remove a gene, a DNA segment encoding the Cas9 enzyme and the guide RNAs responsible for the cut would take the place of the missing gene in the genome. This allowed gene deletions to repeat and propagate until all copies were removed.
Our data indicate that complex viruses arose from smaller and simpler ones.
—Hugo Bisio, Aix-Marseille University
Once they optimized their CRISPR/Cas9 system, the team deleted each gene separately from the Pandoravirus genome and measured the resulting change in virus production, in order to determine how important each gene is to the virus’s lifecycle. They found that essential genes clustered together at one end of the genome and were segregated from nonessential genes at the other end. This level of gene orderliness has not been seen in viruses, according to Bisio. Even bacterial genomes aren’t quite so tidy: While they do group genes with linked functions together into gene clusters known as operons, these tend to be dispersed throughout the genome rather than grouped all together in one spot.
Bisio says the cluster of essential genes may echo a smaller “core genome” of an ancient virus. This genome could have become elongated through multiple rounds of gene duplication that were biased in one direction to produce an additional set of spare nonessential genes. This could explain how modern-day giant viruses came to possess thousands of genes. “Our data indicate that complex viruses arose from smaller and simpler ones,” Bisio tells The Scientist in an email—noting that it will take further research to determine whether that’s true of all giant viruses or just Pandoravirus. Other studies found that some genes in giant viruses were usurped from their amoeba hosts, suggesting gene exchange is another way giant viruses increased in size.
See “New Giant Virus Group Reported”
The team then set their sights on one of the many evolutionary mysteries of Pandoravirus: its lack of a capsid. Small viruses package their genomes into capsids made of viral proteins. While some giant viruses, such as Mimivirus, continue this tradition, others, including Pandoravirus, do not. If giant viruses did indeed evolve from smaller ones, there could be traces of capsid proteins hiding in their genomes. So, the researchers set out to study the function of potential capsid protein remnants in a close cousin of Pandoravirus, the smaller Mollivirus, which can also infect A. castellanii.
See “Another Ancient Giant Virus Discovered”
Researchers have suspected that a Mollivirus protein called ml_347 evolved from a capsid gene based on its gene’s sequence and predicted 3D shape. So, the team investigated its function by deleting the gene using their CRISPR/Cas9 system. They found that the gene is important for Mollivirus assembly, which the authors say is intriguing given its possible capsid ancestry. It’s possible that, as capsids were lost in giant virus evolution, obsolete capsid genes were adapted for new assembly functions.
Frederik Schulz, an evolutionary biologist with the DOE Joint Genome Institute in California who wasn’t involved with the study but who has worked with Chantal Abergel in the past, tells The Scientist that the findings align with recent discoveries. “There was a debate for a long time [about] how giant virus[es] evolved,” he says. “The working hypothesis in the previous years was that they evolved from smaller viruses, and that’s exactly what Chantal and her team could show and confirm using their CRISPR/Cas9 gene-editing approach.”
Schulz notes that it will be exciting to see the CRISPR/Cas9 technology introduced into other host species, such as algae, which would allow researchers to expand research into a greater variety of giant viruses. He also points out that the system only works for viruses that replicate in a host cell’s nucleus, while most giant viruses replicate in cytoplasmic structures called viral factories, which the Cas9 enzyme and guide RNAs can’t penetrate.
Still, Bisio says there’s much left to discover in Pandoravirus. “[This CRISPR/Cas9 technology is] a goldmine to find new functions,” he says—one that he and his colleagues are eager to employ to tease apart what all the virus’s genes do.